Recent News

- News
Genome-wide association analyses identify 95 risk loci and provide insights into the neurobiology of post-traumatic stress disorder
Global PTSD Genetics Partnership achieves new milestone

- News
New meta-analysis builds upon PTSD genetics research, providing a foundation for the development of precision therapies
- News
Driving Progress: TBI Action Alliance Launches Multi-Pronged Initiative to Speed Development of Diagnostic and Therapeutic Solutions for Traumatic Brain Injury
- news
CVB President Nicole Harmon, PhD Discusses How Traumatic Brain Injury Has Impacted Her Family
News
Genome-wide association analyses identify 95 risk loci and provide insights into the neurobiology of post-traumatic stress disorder
Global PTSD Genetics Partnership achieves new milestone
Learn More
News
New meta-analysis builds upon PTSD genetics research, providing a foundation for the development of precision therapies
Learn More
News
Driving Progress: TBI Action Alliance Launches Multi-Pronged Initiative to Speed Development of Diagnostic and Therapeutic Solutions for Traumatic Brain Injury
Learn More
news
CVB President Nicole Harmon, PhD Discusses How Traumatic Brain Injury Has Impacted Her Family
Learn MoreEvents

- Event
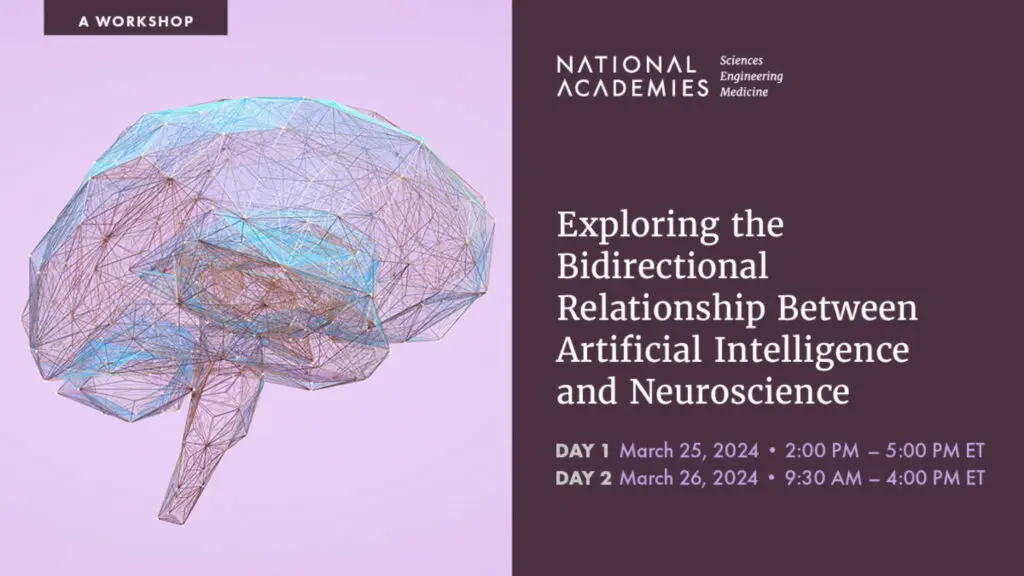
National Academies Workshop: Exploring the Bidirectional Relationship between Artificial Intelligence and Neuroscience
The intersection of artificial intelligence (AI) and neuroscience holds tremendous promise for advancements in brain health. To explore this complex bidirectional relationship, Cohen Veterans Bioscience (CVB) is proud to have

- Event
First Responders Town Hall Event
You're invited to attend an interactive town hall style event in support of the first responder community.

- Event
22 Jumps: February 3, 2024 at Camelback Mountain, Arizona
Three BASE jumpers are BASE jumping 22 times in a single day from Arizona's Camelback Mountain on February 3rd, 2024, seeking to raise more than $22,000 in honor of the

- Event
Webinar – Data-Driven Solutions for Advancing Brain Health Research: Optimizing Research Workflows
The goal of this webinar is to promote awareness of the advances and challenges in data management and sharing and to facilitate conversation among the TBI clinical research community where

- Event
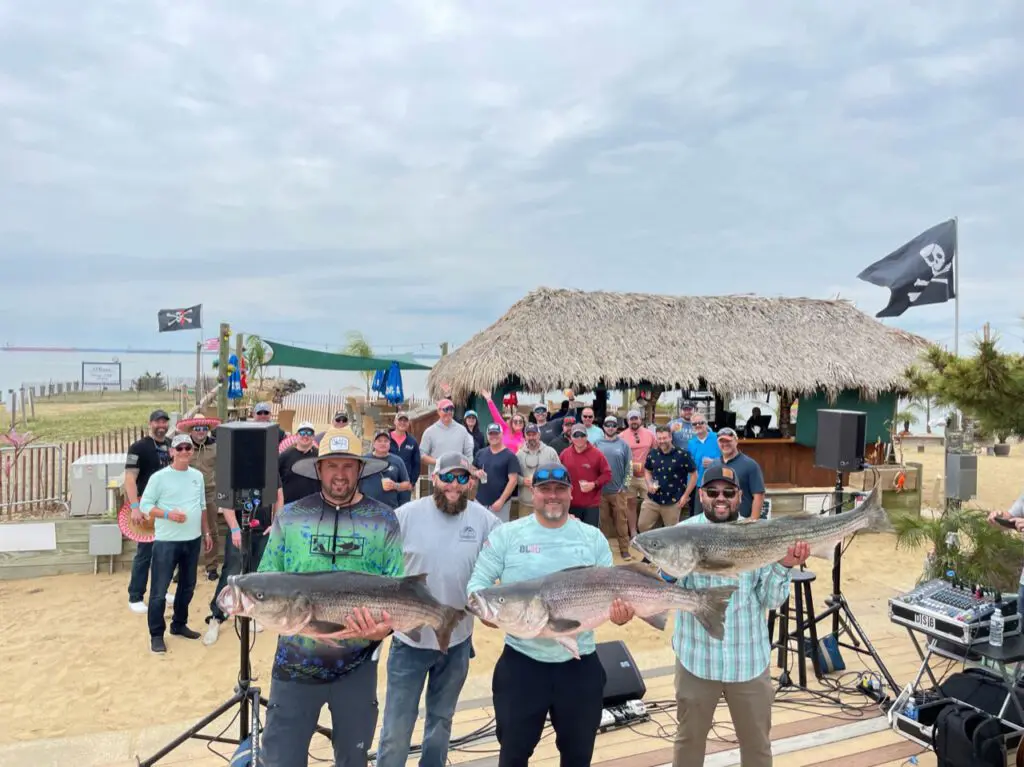
2023 Memorial Rock N’ 4 Ryan Rockfish Tournament to support traumatic brain injury research
The second annual Memorial Rock N' 4 Ryan Rockfish Tournament will be held on October 19, 2023, in honor of Ryan Larkin, a decorated Navy SEAL who served four tours

- Event
22 Jumps: October 21, 2023 in Fayetteville, West Virginia
22 Jumps is excited to BASE jump at the New River Gorge Bridge in Fayetteville, West Virginia on October 21, 2023.

- Event
22 Jumps: Memorial Day Weekend 2023, Twin Falls, Idaho
On Saturday, May 27, 2023, watch as four jumpers leap from the Perrine Bridge in Twin Falls, Idaho 22 times each in a single day — 22 symbolic of the

- Event
22 Jumps: October 15, 2022 in Fayetteville, West Virginia
22 Jumps is excited to BASE jump at the New River Gorge Bridge in Fayetteville, West Virginia on October 15, 2022. 8 jumpers will be doing a combined 22 jumps
Event
National Academies Workshop: Exploring the Bidirectional Relationship between Artificial Intelligence and Neuroscience
The intersection of artificial intelligence (AI) and neuroscience holds tremendous promise for advancements in brain health. To explore this complex bidirectional relationship, Cohen Veterans Bioscience (CVB) is proud to have our Founder, Magali Haas, MD, PhD co-chair the upcoming workshop, hosted by the National Academies of Sciences, Engineering, and Medicine, “Exploring the Bidirectional Relationship Between Artificial Intelligence and Neuroscience.” CVB has been a long-term member of the NASEM Neuroscience Forum.
Learn More
Event
First Responders Town Hall Event
You're invited to attend an interactive town hall style event in support of the first responder community.
Learn More
Event
22 Jumps: February 3, 2024 at Camelback Mountain, Arizona
Three BASE jumpers are BASE jumping 22 times in a single day from Arizona's Camelback Mountain on February 3rd, 2024, seeking to raise more than $22,000 in honor of the 22 Veterans and service members who lose their battle to suicide each day.
Learn More
Event
Webinar – Data-Driven Solutions for Advancing Brain Health Research: Optimizing Research Workflows
The goal of this webinar is to promote awareness of the advances and challenges in data management and sharing and to facilitate conversation among the TBI clinical research community where there is a shared mission for improved outcomes and an overall desire to accelerate discovery in support of brain health.
Learn More
Event
2023 Memorial Rock N’ 4 Ryan Rockfish Tournament to support traumatic brain injury research
The second annual Memorial Rock N' 4 Ryan Rockfish Tournament will be held on October 19, 2023, in honor of Ryan Larkin, a decorated Navy SEAL who served four tours of duty during more than a decade of service.
Learn More
Event
22 Jumps: October 21, 2023 in Fayetteville, West Virginia
22 Jumps is excited to BASE jump at the New River Gorge Bridge in Fayetteville, West Virginia on October 21, 2023.
Learn More
Event
22 Jumps: Memorial Day Weekend 2023, Twin Falls, Idaho
On Saturday, May 27, 2023, watch as four jumpers leap from the Perrine Bridge in Twin Falls, Idaho 22 times each in a single day — 22 symbolic of the estimated 22 servicemembers and Veterans who take their lives each day.
Learn More
Event
22 Jumps: October 15, 2022 in Fayetteville, West Virginia
22 Jumps is excited to BASE jump at the New River Gorge Bridge in Fayetteville, West Virginia on October 15, 2022. 8 jumpers will be doing a combined 22 jumps throughout the day.
Learn More
Featured Articles
- Blog
Discussing the Normative Neuroimaging Library with Dr. James Stone
James R. Stone, MD, PhD is the scientific principal investigator for the Normative Neuroimaging Library, and Associate Professor and Vice Chair of Research, UVA Department of Radiology and Medical Imaging.
- Blog
Thoughts on COVID-19 from Frank Larkin, Chair of the Veterans Advisory Council
"We will get through this, but not without experiencing a variable degree of dysfunction, inconvenience and pain. The greatest threat, beyond the unknown" surrounding this epidemic, is FEAR. Fear can

- Blog
A Navy SEAL Talks Commitment and Care
Brian Losey has spent his entire career—some 33 years—serving our country. In August 2016, he retired as Rear Admiral of the U.S. Navy where he led the Special Warfare Command.

- Blog
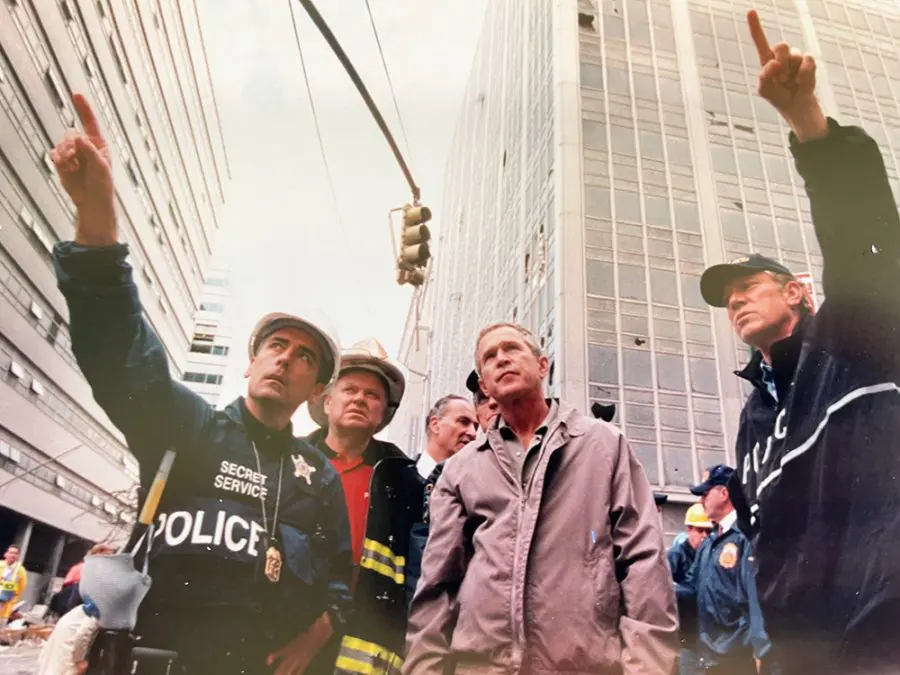
Reflections On The Impact Of 9/11 From A Secret Service Agent Who Was There – And Whose Family Was Forever Changed
Experience Sept. 11, 2001 from Frank Larkin, US Secret Service senior supervisor/special agent assigned to the New York Field Office located in Building #7 of the New York City World

- Blog
How to Maintain Your Mental Health
Research suggests long-term stress can have a negative impact on the brain. When chronic stress causes our brains to remain in a fight or flight state, we can experience a
- Blog
Understanding How Suicide Relates to PTSD and TBI
September is Suicide Prevention Awareness Month, and September 10th is World Suicide Prevention Day. Every year, more than 41,000 people die by suicide in the United States. During September, you

- Blog
Q&A Spotlight with Lee Lancashire, PhD, on PTSD Data Modeling
Chief Information Officer Dr. Lee Lancashire discusses how CVB's approach to data modeling is helping to advance research for trauma-related brain disorders, such as PTSD and TBI.

- Blog
Understanding Sexual Assault and PTSD
The goal of Sexual Assault Awareness Month (SAAM) is to increase awareness of the prevalence of sexual violence and to provide education to individuals and communities about prevention. Sexual violence
Blog
Discussing the Normative Neuroimaging Library with Dr. James Stone
James R. Stone, MD, PhD is the scientific principal investigator for the Normative Neuroimaging Library, and Associate Professor and Vice Chair of Research, UVA Department of Radiology and Medical Imaging.
Learn More
Blog
Thoughts on COVID-19 from Frank Larkin, Chair of the Veterans Advisory Council
"We will get through this, but not without experiencing a variable degree of dysfunction, inconvenience and pain. The greatest threat, beyond the unknown" surrounding this epidemic, is FEAR. Fear can be crippling and can take you to a very dark place of despair and hopelessness. Fear can also tune your senses and personal resolve to overcome a threat."
Learn More
Blog
A Navy SEAL Talks Commitment and Care
Brian Losey has spent his entire career—some 33 years—serving our country. In August 2016, he retired as Rear Admiral of the U.S. Navy where he led the Special Warfare Command.
Learn More
Blog
Reflections On The Impact Of 9/11 From A Secret Service Agent Who Was There – And Whose Family Was Forever Changed
Experience Sept. 11, 2001 from Frank Larkin, US Secret Service senior supervisor/special agent assigned to the New York Field Office located in Building #7 of the New York City World Trade Center (WTC) complex.
Learn More
Blog
How to Maintain Your Mental Health
Research suggests long-term stress can have a negative impact on the brain. When chronic stress causes our brains to remain in a fight or flight state, we can experience a host of negative effects on both our brains and bodies
Learn More
Blog
Understanding How Suicide Relates to PTSD and TBI
September is Suicide Prevention Awareness Month, and September 10th is World Suicide Prevention Day. Every year, more than 41,000 people die by suicide in the United States. During September, you can help combat stigma and raise awareness about suicide and prevention, connect with people you know who are impacted, and share resources to help those at risk.
Learn More
Blog
Q&A Spotlight with Lee Lancashire, PhD, on PTSD Data Modeling
Chief Information Officer Dr. Lee Lancashire discusses how CVB's approach to data modeling is helping to advance research for trauma-related brain disorders, such as PTSD and TBI.
Learn More
Blog
Understanding Sexual Assault and PTSD
The goal of Sexual Assault Awareness Month (SAAM) is to increase awareness of the prevalence of sexual violence and to provide education to individuals and communities about prevention. Sexual violence often has long-term effects on victims, including Post-Traumatic Stress Disorder (PTSD) and suicidality.
Learn More















